
A transposable element is a nucleic acid sequence in DNA that can change its position within a genome, sometimes creating or reversing mutations and altering the cell's genetic identity and genome size. Transposition often results in duplication of the same genetic material. In the human genome, L1 and Alu elements are two examples. Barbara McClintock's discovery of them earned her a Nobel Prize in 1983. Its importance in personalized medicine is becoming increasingly relevant, as well as gaining more attention in data analytics given the difficulty of analysis in very high dimensional spaces.
Small RNA (sRNA) are polymeric RNA molecules that are less than 200 nucleotides in length, and are usually non-coding. RNA silencing is often a function of these molecules, with the most common and well-studied example being RNA interference (RNAi), in which endogenously expressed microRNA (miRNA) or exogenously derived small interfering RNA (siRNA) induces the degradation of complementary messenger RNA. Other classes of small RNA have been identified, including piwi-interacting RNA (piRNA) and its subspecies repeat associated small interfering RNA (rasiRNA). Small RNA "is unable to induce RNAi alone, and to accomplish the task it must form the core of the RNA–protein complex termed the RNA-induced silencing complex (RISC), specifically with Argonaute protein".
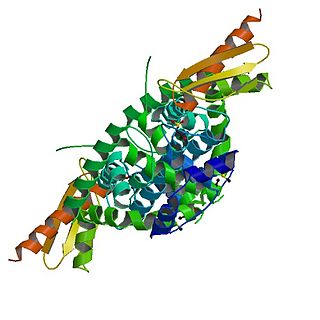
Dicer, also known as endoribonuclease Dicer or helicase with RNase motif, is an enzyme that in humans is encoded by the DICER1 gene. Being part of the RNase III family, Dicer cleaves double-stranded RNA (dsRNA) and pre-microRNA (pre-miRNA) into short double-stranded RNA fragments called small interfering RNA and microRNA, respectively. These fragments are approximately 20–25 base pairs long with a two-base overhang on the 3′-end. Dicer facilitates the activation of the RNA-induced silencing complex (RISC), which is essential for RNA interference. RISC has a catalytic component Argonaute, which is an endonuclease capable of degrading messenger RNA (mRNA).
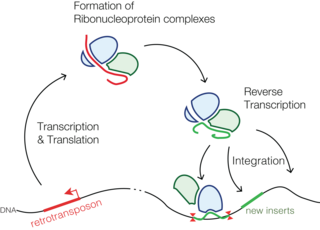
Retrotransposons are a type of genetic component that copy and paste themselves into different genomic locations (transposon) by converting RNA back into DNA through the reverse transcription process using an RNA transposition intermediate.
The RNA-induced silencing complex, or RISC, is a multiprotein complex, specifically a ribonucleoprotein, which functions in gene silencing via a variety of pathways at the transcriptional and translational levels. Using single-stranded RNA (ssRNA) fragments, such as microRNA (miRNA), or double-stranded small interfering RNA (siRNA), the complex functions as a key tool in gene regulation. The single strand of RNA acts as a template for RISC to recognize complementary messenger RNA (mRNA) transcript. Once found, one of the proteins in RISC, Argonaute, activates and cleaves the mRNA. This process is called RNA interference (RNAi) and it is found in many eukaryotes; it is a key process in defense against viral infections, as it is triggered by the presence of double-stranded RNA (dsRNA).

Hop, occasionally written HOP, is an abbreviation for Hsp70-Hsp90 Organizing Protein. It functions as a co-chaperone which reversibly links together the protein chaperones Hsp70 and Hsp90.
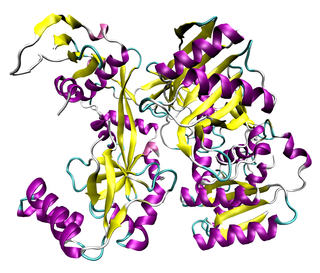
The Argonaute protein family, first discovered for its evolutionarily conserved stem cell function, plays a central role in RNA silencing processes as essential components of the RNA-induced silencing complex (RISC). RISC is responsible for the gene silencing phenomenon known as RNA interference (RNAi). Argonaute proteins bind different classes of small non-coding RNAs, including microRNAs (miRNAs), small interfering RNAs (siRNAs) and Piwi-interacting RNAs (piRNAs). Small RNAs guide Argonaute proteins to their specific targets through sequence complementarity, which then leads to mRNA cleavage, translation inhibition, and/or the initiation of mRNA decay.
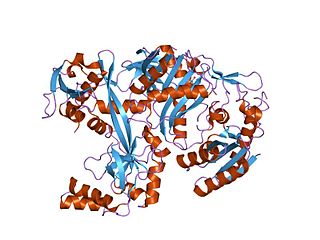
Piwi genes were identified as regulatory proteins responsible for stem cell and germ cell differentiation. Piwi is an abbreviation of P-elementInduced WImpy testis in Drosophila. Piwi proteins are highly conserved RNA-binding proteins and are present in both plants and animals. Piwi proteins belong to the Argonaute/Piwi family and have been classified as nuclear proteins. Studies on Drosophila have also indicated that Piwi proteins have no slicer activity conferred by the presence of the Piwi domain. In addition, Piwi associates with heterochromatin protein 1, an epigenetic modifier, and piRNA-complementary sequences. These are indications of the role Piwi plays in epigenetic regulation. Piwi proteins are also thought to control the biogenesis of piRNA as many Piwi-like proteins contain slicer activity which would allow Piwi proteins to process precursor piRNA into mature piRNA.
RNA silencing or RNA interference refers to a family of gene silencing effects by which gene expression is negatively regulated by non-coding RNAs such as microRNAs. RNA silencing may also be defined as sequence-specific regulation of gene expression triggered by double-stranded RNA (dsRNA). RNA silencing mechanisms are conserved among most eukaryotes. The most common and well-studied example is RNA interference (RNAi), in which endogenously expressed microRNA (miRNA) or exogenously derived small interfering RNA (siRNA) induces the degradation of complementary messenger RNA. Other classes of small RNA have been identified, including piwi-interacting RNA (piRNA) and its subspecies repeat associated small interfering RNA (rasiRNA).
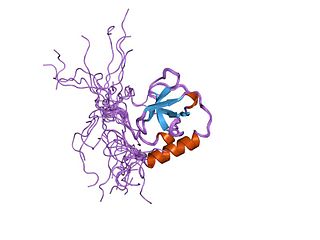
In molecular biology, a Tudor domain is a conserved protein structural domain originally identified in the Tudor protein encoded in Drosophila. The Tudor gene was found in a Drosophila screen for maternal factors that regulate embryonic development or fertility. Mutations here are lethal for offspring, inspiring the name Tudor, as a reference to the Tudor King Henry VIII and the several miscarriages experienced by his wives.
Nuage are Drosophila melanogaster germline granules. Nuage are the hallmark of Drosophila melanogaster germline cells, which have an electron-dense perinuclear structure and can silence the selfish genetic elements in Drosophila melanogaster. The term 'Nuage' comes from the French word for 'cloud', as they appear as nebulous electron-dense bodies by electron microscopy. They are found in nurse cells of the developing Drosophila melanogaster egg chamber and are composed of various types of proteins, including RNA-helicases, Tudor domain proteins, Piwi-clade Argonaute proteins, in addition to a PRMT5 methylosome composed of Capsuléen and its co-factor, Valois (MEP50).
Transposon silencing is a form of transcriptional gene silencing targeting transposons. Transcriptional gene silencing is a product of histone modifications that prevent the transcription of a particular area of DNA. Transcriptional silencing of transposons is crucial to the maintenance of a genome. The “jumping” of transposons generates genomic instability and can cause extremely deleterious mutations. Transposable element insertions have been linked to many diseases including hemophilia, severe combined immunodeficiency, and predisposition to cancer. The silencing of transposons is therefore extremely critical in the germline in order to stop transposon mutations from developing and being passed on to the next generation. Additionally, these epigenetic defenses against transposons can be heritable. Studies in Drosophila, Arabidopsis thaliana, and mice all indicate that small interfering RNAs are responsible for transposon silencing. In animals, these siRNAS and piRNAs are most active in the gonads.
RDE-1 (RNAi-DEfective 1) is a primary Argonaute protein required for RNA-mediated interference (RNAi) in Caenorhabditis elegans. The rde-1 gene locus was first characterized in C. elegans mutants resistant to RNAi, and is a member of a highly conserved Piwi gene family that includes plant, Drosophila, and vertebrate homologs.
The gene Maelstrom, Mael, creates a protein, which was first located in Drosophila melanogaster in the nuage perinuclear structure and has functionality analogous to the spindle, spn, gene class. Its mammalian homolog is MAEL.

Transposable elements are pieces of genetic material that are capable of splicing themselves into a host genome and then self propagating throughout the genome, much like a virus. Retrotransposons are a subset of transposable elements that use an RNA intermediate and reverse transcribe themselves into the genome. Retrotransposon proliferation may lead to insertional mutagenesis, disrupt the process of DNA repair, or cause errors during chromosomal crossover, and so it is advantageous for an organism to possess the means to suppress or "silence" retrotransposon activity.
Ruth Lehmann is a developmental and cell biologist. She is the Director of the Whitehead Institute for Biomedical Research. She previously was affiliated with the New York University School of Medicine, where she was the Director of the Skirball Institute of Biomolecular Medicine, the Laura and Isaac Perlmutter Professor of Cell Biology, and the Chair of the Department of Cell Biology. Her research focuses on germ cells and embryogenesis.
piwi-interacting RNA (piRNA) belongs to the small RNA class found in eukaryotic organisms, their major role is to regulate the expression of genes by the mRNA degradation or silencing. After the association with argonaute protein family, these short RNA guide the RNA-induced silencing complex (RISC) to their target by sequence complementarity. piRNAs have approximately from 26 to 31 nucleotides and are found in almost all metazoans.
Haifan Lin is a Chinese-born American stem cell biologist. He is the Eugene Higgins Chair Professor of Cell Biology at Yale University and the founding Director of the Yale Stem Cell Center. He previously founded and directed the Stem Cell Research Program at Duke University. Recognized for his significant contributions to stem cell research, he was elected to the US National Academy of Sciences and American Academy of Arts and Sciences in 2018.
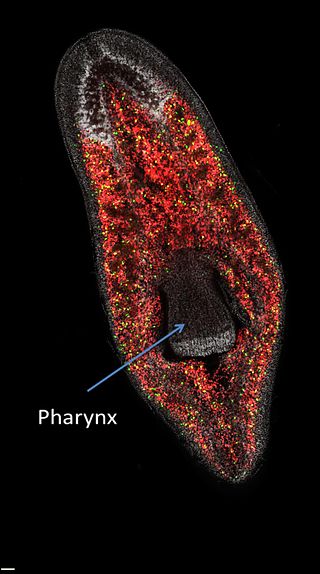
Neoblasts (ˈniːəʊˌblæst) are adult stem cells found in planarian flatworms. They are the only dividing planarian cells, and they produce all cell types, including the germline. Neoblasts are abundant in the planarian parenchyma, and comprise up to 30 percent of all cells. Following injury, neoblasts rapidly divide and generate new cells, which allows planarians to regenerate any missing tissue.









