
Bacillus thuringiensis is a gram-positive, soil-dwelling bacterium, the most commonly used biological pesticide worldwide. B. thuringiensis also occurs naturally in the gut of caterpillars of various types of moths and butterflies, as well on leaf surfaces, aquatic environments, animal feces, insect-rich environments, flour mills and grain-storage facilities. It has also been observed to parasitize moths such as Cadra calidella—in laboratory experiments working with C. calidella, many of the moths were diseased due to this parasite.

A biofilm is a syntrophic community of microorganisms in which cells stick to each other and often also to a surface. These adherent cells become embedded within a slimy extracellular matrix that is composed of extracellular polymeric substances (EPSs). The cells within the biofilm produce the EPS components, which are typically a polymeric combination of extracellular polysaccharides, proteins, lipids and DNA. Because they have a three-dimensional structure and represent a community lifestyle for microorganisms, they have been metaphorically described as "cities for microbes".
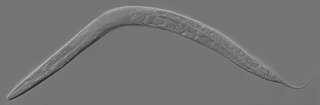
Caenorhabditis elegans is a free-living transparent nematode about 1 mm in length that lives in temperate soil environments. It is the type species of its genus. The name is a blend of the Greek caeno- (recent), rhabditis (rod-like) and Latin elegans (elegant). In 1900, Maupas initially named it Rhabditides elegans. Osche placed it in the subgenus Caenorhabditis in 1952, and in 1955, Dougherty raised Caenorhabditis to the status of genus.

Lipopolysaccharide, now more commonly known as Endotoxin, is a collective term for components of the outermost membrane of cell envelope of Gram-negative bacteria, such as E. coli and Salmonella. with a common structural architecture. Lipopolysaccharides (LPS) are large molecules consisting of three parts: an outer core polysaccharide termed the O-antigen, an inner core oligosaccharide and Lipid A, all covalently linked. In current terminology, the term endotoxin is often used synonymously with LPS, although there are a few endotoxins that are not related to LPS, such as the so-called delta endotoxin proteins produced by Bacillus thuringiensis.

Pseudomonas aeruginosa is a common encapsulated, Gram-negative, aerobic–facultatively anaerobic, rod-shaped bacterium that can cause disease in plants and animals, including humans. A species of considerable medical importance, P. aeruginosa is a multidrug resistant pathogen recognized for its ubiquity, its intrinsically advanced antibiotic resistance mechanisms, and its association with serious illnesses – hospital-acquired infections such as ventilator-associated pneumonia and various sepsis syndromes. P. aeruginosa is able to selectively inhibit various antibiotics from penetrating its outer membrane - and has high resistance to several antibiotics. According to the World Health Organization P. aeruginosa poses one of the greatest threats to humans in terms of antibiotic resistance.
Adhesins are cell-surface components or appendages of bacteria that facilitate adhesion or adherence to other cells or to surfaces, usually in the host they are infecting or living in. Adhesins are a type of virulence factor.
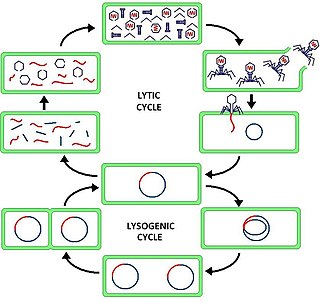
Lysogeny, or the lysogenic cycle, is one of two cycles of viral reproduction. Lysogeny is characterized by integration of the bacteriophage nucleic acid into the host bacterium's genome or formation of a circular replicon in the bacterial cytoplasm. In this condition the bacterium continues to live and reproduce normally, while the bacteriophage lies in a dormant state in the host cell. The genetic material of the bacteriophage, called a prophage, can be transmitted to daughter cells at each subsequent cell division, and later events can release it, causing proliferation of new phages via the lytic cycle.
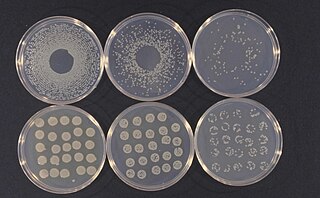
Pseudomonas syringae is a rod-shaped, Gram-negative bacterium with polar flagella. As a plant pathogen, it can infect a wide range of species, and exists as over 50 different pathovars, all of which are available to researchers from international culture collections such as the NCPPB, ICMP, and others.

Pathogenic bacteria are bacteria that can cause disease. This article focuses on the bacteria that are pathogenic to humans. Most species of bacteria are harmless and are often beneficial but others can cause infectious diseases. The number of these pathogenic species in humans is estimated to be fewer than a hundred. By contrast, several thousand species are part of the gut flora present in the digestive tract.
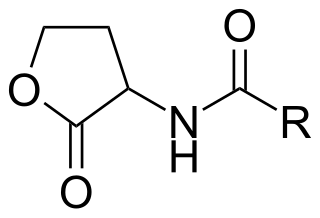
Lactonase (EC 3.1.1.81, acyl-homoserine lactonase; systematic name N-acyl-L-homoserine-lactone lactonohydrolase) is a metalloenzyme, produced by certain species of bacteria, which targets and inactivates acylated homoserine lactones (AHLs). It catalyzes the reaction

Bacillus anthracis is a gram-positive and rod-shaped bacterium that causes anthrax, a deadly disease to livestock and, occasionally, to humans. It is the only permanent (obligate) pathogen within the genus Bacillus. Its infection is a type of zoonosis, as it is transmitted from animals to humans. It was discovered by a German physician Robert Koch in 1876, and became the first bacterium to be experimentally shown as a pathogen. The discovery was also the first scientific evidence for the germ theory of diseases.
Pathogenomics is a field which uses high-throughput screening technology and bioinformatics to study encoded microbe resistance, as well as virulence factors (VFs), which enable a microorganism to infect a host and possibly cause disease. This includes studying genomes of pathogens which cannot be cultured outside of a host. In the past, researchers and medical professionals found it difficult to study and understand pathogenic traits of infectious organisms. With newer technology, pathogen genomes can be identified and sequenced in a much shorter time and at a lower cost, thus improving the ability to diagnose, treat, and even predict and prevent pathogenic infections and disease. It has also allowed researchers to better understand genome evolution events - gene loss, gain, duplication, rearrangement - and how those events impact pathogen resistance and ability to cause disease. This influx of information has created a need for bioinformatics tools and databases to analyze and make the vast amounts of data accessible to researchers, and it has raised ethical questions about the wisdom of reconstructing previously extinct and deadly pathogens in order to better understand virulence.
Host–parasite coevolution is a special case of coevolution, where a host and a parasite continually adapt to each other. This can create an evolutionary arms race between them. A more benign possibility is of an evolutionary trade-off between transmission and virulence in the parasite, as if it kills its host too quickly, the parasite will not be able to reproduce either. Another theory, the Red Queen hypothesis, proposes that since both host and parasite have to keep on evolving to keep up with each other, and since sexual reproduction continually creates new combinations of genes, parasitism favours sexual reproduction in the host.
Jonathan Alan Hodgkin is a British biochemist. He is the Professor of Genetics at the University of Oxford and an emeritus fellow of Keble College, Oxford.
In biology, a pathogen, in the oldest and broadest sense, is any organism or agent that can produce disease. A pathogen may also be referred to as an infectious agent, or simply a germ.
The host–pathogen interaction is defined as how microbes or viruses sustain themselves within host organisms on a molecular, cellular, organismal or population level. This term is most commonly used to refer to disease-causing microorganisms although they may not cause illness in all hosts. Because of this, the definition has been expanded to how known pathogens survive within their host, whether they cause disease or not.

Frederick M Ausubel is an American molecular biologist and professor of genetics at Harvard Medical School in Boston and is the Karl Winnacker Distinguished Investigator in the Department of Molecular Biology, Massachusetts General Hospital, Boston., Massachusetts.
Nematocida parisii is a parasitic species of Microsporidia fungi found in wild isolates of the common nematode, Caenorhabditis elegans. The fungus forms spores and replicates in the intestines before leaving the host.
Worm bagging is a form of vivipary observed in nematodes, namely Caenorhabditis elegans. The process is characterized by eggs hatching within the parent and the larvae proceeding to consume and emerge from the parent.
Pathogen avoidance, also referred to as, parasite avoidance or pathogen disgust, refers to the theory that the disgust response, in humans, is an adaptive system that guides behavior to avoid infection caused by parasites such as viruses, bacteria, fungi, protozoa, helminth worms, arthropods and social parasites. Pathogen avoidance is a psychological mechanism associated with the behavioral immune system. Pathogen avoidance has been discussed as one of the three domains of disgust which also include sexual and moral disgust.














