In the fields of molecular biology and genetics, a genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA. The nuclear genome includes protein-coding genes and non-coding genes, other functional regions of the genome such as regulatory sequences, and often a substantial fraction of junk DNA with no evident function. Almost all eukaryotes have mitochondria and a small mitochondrial genome. Algae and plants also contain chloroplasts with a chloroplast genome.
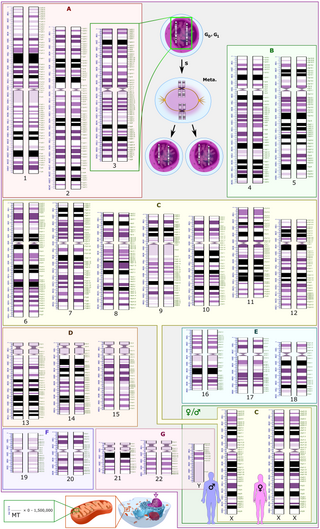
The human genome is a complete set of nucleic acid sequences for humans, encoded as DNA within the 23 chromosome pairs in cell nuclei and in a small DNA molecule found within individual mitochondria. These are usually treated separately as the nuclear genome and the mitochondrial genome. Human genomes include both protein-coding DNA sequences and various types of DNA that does not encode proteins. The latter is a diverse category that includes DNA coding for non-translated RNA, such as that for ribosomal RNA, transfer RNA, ribozymes, small nuclear RNAs, and several types of regulatory RNAs. It also includes promoters and their associated gene-regulatory elements, DNA playing structural and replicatory roles, such as scaffolding regions, telomeres, centromeres, and origins of replication, plus large numbers of transposable elements, inserted viral DNA, non-functional pseudogenes and simple, highly repetitive sequences. Introns make up a large percentage of non-coding DNA. Some of this non-coding DNA is non-functional junk DNA, such as pseudogenes, but there is no firm consensus on the total amount of junk DNA.

BGI Group, formerly Beijing Genomics Institute, is a Chinese genomics company with headquarters in Yantian, Shenzhen. The company was originally formed in 1999 as a genetics research center to participate in the Human Genome Project. It also sequences the genomes of other animals, plants and microorganisms.
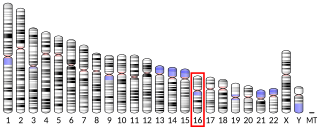
Growth factor, augmenter of liver regeneration , also known as GFER, or Hepatopoietin is a protein which in humans is encoded by the GFER gene. This gene is also known as essential for respiration and vegatative growth, augmenter of liver regeneration, and growth factor of Erv1-like/Hepatic regenerative stimulation substance.

Integral membrane protein 2C is a protein that in humans is encoded by the ITM2C gene.

Ubiquitin-associated protein 1 is a protein that in humans is encoded by the UBAP1 gene.

AT-rich interactive domain-containing protein 2 (ARID2) is a protein that in humans is encoded by the ARID2 gene.

Biogenesis of lysosome-related organelles complex 1 subunit 2 is a protein that in humans is encoded by the BLOC1S2 gene.

Spartan (SPRTN) is a protein that in humans is encoded by the SPRTN gene. It is involved in DNA repair. Ruijs-Aalfs syndrome is an autosomal recessive genetic disorder. Characteristics of this disorder are features of premature aging, chromosome instability and development of hepatocellular carcinoma. Ruijs-Aalfs syndrome arises as a result of mutations in the SPRTN gene that encodes a metalloproteinase employed in the repair of protein-linked DNA breaks.
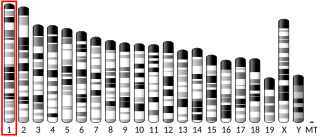
Ribosomal protein S6 kinase delta-1 is an enzyme that in humans is encoded by the RPS6KC1 gene.
39S ribosomal protein L32, mitochondrial is a protein that in humans is encoded by the MRPL32 gene.

Transmembrane protein 8B is a protein that in humans is encoded by the TMEM8B gene. It encodes for a transmembrane protein that is 338 amino acids long, and is located on human chromosome 9. Aliases associated with this gene include C9orf127, NAG-5, and NGX61.

Whole genome sequencing (WGS) is the process of determining the entirety, or nearly the entirety, of the DNA sequence of an organism's genome at a single time. This entails sequencing all of an organism's chromosomal DNA as well as DNA contained in the mitochondria and, for plants, in the chloroplast.
Complete Genomics is a life sciences company that has developed and commercialized a DNA sequencing platform for human genome sequencing and analysis. This solution combines the company's proprietary human genome sequencing technology with its informatics and data management software to provide finished variant reports and assemblies at Complete Genomics’ commercial genome center in Mountain View, California.
SOAP is a suite of bioinformatics software tools from the BGI Bioinformatics department enabling the assembly, alignment, and analysis of next generation DNA sequencing data. It is particularly suited to short read sequencing data.

Solute carrier family 52, member 3, formerly known as chromosome 20 open reading frame 54 and riboflavin transporter 2, is a protein that in humans is encoded by the SLC52A3 gene.

Genome evolution is the process by which a genome changes in structure (sequence) or size over time. The study of genome evolution involves multiple fields such as structural analysis of the genome, the study of genomic parasites, gene and ancient genome duplications, polyploidy, and comparative genomics. Genome evolution is a constantly changing and evolving field due to the steadily growing number of sequenced genomes, both prokaryotic and eukaryotic, available to the scientific community and the public at large.
Transcriptomics technologies are the techniques used to study an organism's transcriptome, the sum of all of its RNA transcripts. The information content of an organism is recorded in the DNA of its genome and expressed through transcription. Here, mRNA serves as a transient intermediary molecule in the information network, whilst non-coding RNAs perform additional diverse functions. A transcriptome captures a snapshot in time of the total transcripts present in a cell. Transcriptomics technologies provide a broad account of which cellular processes are active and which are dormant. A major challenge in molecular biology is to understand how a single genome gives rise to a variety of cells. Another is how gene expression is regulated.
















