
A biofilm is a syntrophic community of microorganisms in which cells stick to each other and often also to a surface. These adherent cells become embedded within a slimy extracellular matrix that is composed of extracellular polymeric substances (EPSs). The cells within the biofilm produce the EPS components, which are typically a polymeric combination of extracellular polysaccharides, proteins, lipids and DNA. Because they have a three-dimensional structure and represent a community lifestyle for microorganisms, they have been metaphorically described as "cities for microbes".
In biology, quorum sensing or quorum signaling (QS) is the process of cell-to-cell communication that allows bacteria to detect and respond to cell population density by gene regulation, typically as a means of acclimating to environmental disadvantages.

Pseudomonas aeruginosa is a common encapsulated, Gram-negative, aerobic–facultatively anaerobic, rod-shaped bacterium that can cause disease in plants and animals, including humans. A species of considerable medical importance, P. aeruginosa is a multidrug resistant pathogen recognized for its ubiquity, its intrinsically advanced antibiotic resistance mechanisms, and its association with serious illnesses – hospital-acquired infections such as ventilator-associated pneumonia and various sepsis syndromes. P. aeruginosa is able to selectively inhibit various antibiotics from penetrating its outer membrane - and has high resistance to several antibiotics. According to the World Health Organization P. aeruginosa poses one of the greatest threats to humans in terms of antibiotic resistance.
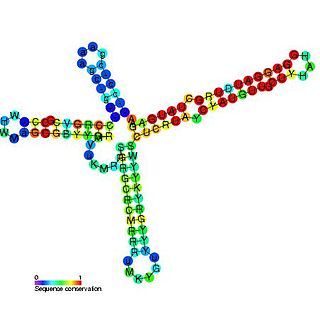
The repression of heat shock gene expression (ROSE) element is an RNA element found in the 5' UTR of some heat shock protein's mRNAs. The ROSE element is an RNA thermometer that negatively regulates heat shock gene expression. The secondary structure is thought to be altered by temperature, thus it is an RNA thermometer. This structure blocks access to the ribosome binding site at normal temperatures. During heat shock however, the structure changes freeing the ribosome binding site and allowing expression to occur.
Biosurfactant usually refers to surfactants of microbial origin. Most of the biosurfactants produced by microbes are synthesized extracellularly and many microbes are known to produce biosurfactants in large relative quantities. Some are of commercial interest. As a secondary metabolite of microorganisms, biosurfactants can be processed by the cultivation of biosurfactant producing microorganisms in the stationary phase on many sorts of low-priced substrates like biochar, plant oils, carbohydrates, wastes, etc. High-level production of biosurfactants can be controlled by regulation of environmental factors and growth circumstances.

Swarming motility is a rapid and coordinated translocation of a bacterial population across solid or semi-solid surfaces, and is an example of bacterial multicellularity and swarm behaviour. Swarming motility was first reported by Jorgen Henrichsen and has been mostly studied in genus Serratia, Salmonella, Aeromonas, Bacillus, Yersinia, Pseudomonas, Proteus, Vibrio and Escherichia.
In enzymology, a succinylglutamate-semialdehyde dehydrogenase (EC 1.2.1.71) is an enzyme that catalyzes the chemical reaction
In biology, an autoinducer is a signaling molecule that enables detection and response to changes in the population density of bacterial cells. Synthesized when a bacterium reproduces, autoinducers pass outside the bacterium and into the surrounding medium. They are a key component of the phenomenon of quorum sensing: as the density of quorum-sensing bacterial cells increases, so does the concentration of the autoinducer. A bacterium’s detection of an autoinducer above some minimum threshold triggers altered gene expression.
In enzymology, an arginine N-succinyltransferase (EC 2.3.1.109) is an enzyme that catalyzes the chemical reaction
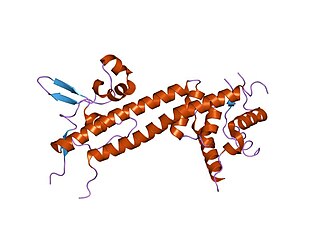
In molecular biology, LcrV is a protein found in Yersinia pestis and several other bacterial species. It forms part of the Yersinia pestis virulence protein factors that also includes all Yops, or Yersinia outer protein, but the name has been kept out of convention. LcrV's main function is not actually known, but it is essential for the production of other Yops.
Roberto Kolter is Professor of Microbiology, Emeritus at Harvard Medical School, an author, and past president of the American Society for Microbiology. Kolter has been a professor at Harvard Medical School since 1983 and was Co-director of Harvard's Microbial Sciences Initiative from 2003-2018. During the 35-year term of the Kolter laboratory from 1983 to 2018, more than 130 graduate student and postdoctoral trainees explored an eclectic mix of topics gravitating around the study of microbes. Kolter is a fellow of the American Association for the Advancement of Science and of the American Academy of Microbiology.
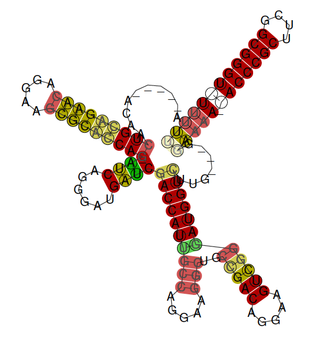
The rsmX gene is part of the Rsm/Csr family of non-coding RNAs (ncRNAs). Members of the Rsm/Csr family are present in a diverse range of bacteria, including Escherichia coli, Erwinia, Salmonella, Vibrio and Pseudomonas. These ncRNAs act by sequestering translational repressor proteins, called RsmA, activating expression of downstream genes that would normally be blocked by the repressors. Sequestering of target proteins is dependent upon exposed GGA motifs in the stem loops of the ncRNAs. Typically, the activated genes are involved in secondary metabolism, biofilm formation and motility.

The PilZ protein family is named after the type IV pilus control protein first identified in Pseudomonas aeruginosa, expressed as part of the pil operon. It has a cytoplasmic location and is essential for type IV fimbrial, or pilus, biogenesis. PilZ is a c-di-GMP binding domain and PilZ domain-containing proteins represent the best studied class of c-di-GMP effectors. C-di-GMP, cyclic diguanosine monophosphate, the second messenger in cells, is widespread in and unique to the bacterial kingdom. Elevated intracellular levels of c-di-GMP generally cause bacteria to change from a motile single-cell state to a sessile, adhesive surface-attached multicellular state called biofilm.
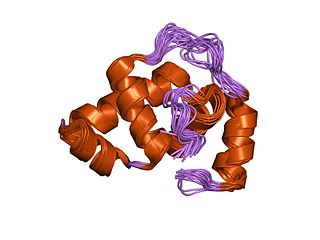
In molecular biology, the LuxR-type DNA-binding HTH domain is a DNA-binding, helix-turn-helix (HTH) domain of about 65 amino acids. It is present in transcription regulators of the LuxR/FixJ family of response regulators. The domain is named after Vibrio fischeri luxR, a transcriptional activator for quorum-sensing control of luminescence. LuxR-type HTH domain proteins occur in a variety of organisms. The DNA-binding HTH domain is usually located in the C-terminal region of the protein; the N-terminal region often containing an autoinducer-binding domain or a response regulatory domain. Most luxR-type regulators act as transcription activators, but some can be repressors or have a dual role for different sites. LuxR-type HTH regulators control a wide variety of activities in various biological processes.
In molecular biology, YadA is a protein domain which is short for Yersinia adhesin A. These proteins have strong sequence and structural homology, particularly at their C-terminal end. The function is to promote their pathogenicity and virulence in host cells, though cell adhesion. YadA is found in three pathogenic species of Yersinia, Y. pestis,Y. pseudotuberculosis, and Y. enterocolitica. The YadA domain is encoded for by a virulence plasmid in Yersinia, which encodes a type-III secretion (T3S) system consisting of the Ysc injectisome and the Yop effectors.
A sophorolipid is a surface-active glycolipid compound that can be synthesized by a selected number of non-pathogenic yeast species. They are potential bio-surfactants due to their biodegradability and low eco-toxicity.

The Catabolite repression control (Crc) protein participates in suppressing expression of several genes involved in utilization of carbon sources in Pseudomonas bacteria. Presence of organic acids triggers activation of Crc and in conjunction with the Hfq protein genes that metabolize a given carbon source are downregulated until another more favorable carbon source is depleted. Crc-mediated regulation impact processes such as biofilm formation, virulence and antibiotic susceptibility.

NrsZ is a bacterial small RNA found in the opportunistic pathogen Pseudomonas aeruginosa PAO1. Its transcription is induced during nitrogen limitation by the NtrB/C two-component system together with the alternative sigma factor RpoN. NrsZ by activating rhlA positively regulates the production of rhamnolipid surfactants needed for swarming motility.
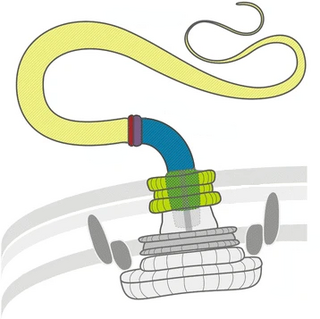
Twitching motility is a form of crawling bacterial motility used to move over surfaces. Twitching is mediated by the activity of hair-like filaments called type IV pili which extend from the cell's exterior, bind to surrounding solid substrates, and retract, pulling the cell forwards in a manner similar to the action of a grappling hook. The name twitching motility is derived from the characteristic jerky and irregular motions of individual cells when viewed under the microscope. It has been observed in many bacterial species, but is most well studied in Pseudomonas aeruginosa, Neisseria gonorrhoeae and Myxococcus xanthus. Active movement mediated by the twitching system has been shown to be an important component of the pathogenic mechanisms of several species.

The molecule 2-heptyl-3-hydroxy-4-quinolone, also named the Pseudomonas quinolone signal (PQS), has been discovered as an intracellular link between the two major quorum sensing systems of P. aeruginosa; the las and rhl systems. These systems together control expression of virulence factors and play a major role in the formation of biofilms in Pseudomonas aeruginosa. P. aeruginosa is a gram-negative bacteria and opportunistic human pathogen that can cause serious and sometimes fatal infections in humans. Similar to other bacterial species, P. aeruginosa uses quorum sensing (QS) systems to communicate between cells in a population. This allows coordination of gene expression in a population based on changing cell densities, abundance of nutrients, and other environmental factors.













