| MLH3 | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | MLH3 , HNPCC7, mutL homolog 3 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | OMIM: 604395 MGI: 1353455 HomoloGene: 91153 GeneCards: MLH3 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
DNA mismatch repair protein Mlh3 is a protein that in humans is encoded by the MLH3 gene. [5] [6]
| MLH3 | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | MLH3 , HNPCC7, mutL homolog 3 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | OMIM: 604395 MGI: 1353455 HomoloGene: 91153 GeneCards: MLH3 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
DNA mismatch repair protein Mlh3 is a protein that in humans is encoded by the MLH3 gene. [5] [6]
This gene is a member of the MutL-homolog (MLH) family of DNA mismatch repair (MMR) genes. MLH genes are implicated in maintaining genomic integrity during DNA replication and after meiotic recombination. The protein encoded by this gene functions as a heterodimer with other family members. Somatic mutations in this gene frequently occur in tumors exhibiting microsatellite instability, and germline mutations have been linked to hereditary nonpolyposis colorectal cancer type 7 (HNPCC7). Several alternatively spliced transcript variants have been identified, but the full-length nature of only two transcript variants has been determined. [6] Orthologs of human MLH3 have also been studied in other organisms including mouse and the budding yeast Saccharomyces cerevisiae .
In addition to its role in DNA mismatch repair, MLH3 protein is also involved in meiotic crossing over. [7] MLH3 forms a heterodimer with MLH1 that appears to be necessary for mouse oocytes to progress through metaphase II of meiosis. [8]

The MLH1-MLH3 heterodimers promote crossovers. [7] Recombination during meiosis is often initiated by a DNA double-strand break (DSB) as illustrated in the accompanying diagram. During recombination, sections of DNA at the 5' ends of the break are cut away in a process called resection. In the strand invasion step that follows, an overhanging 3' end of the broken DNA molecule then "invades" the DNA of a homologous chromosome that is not broken forming a displacement loop (D-loop). After strand invasion, the further sequence of events may follow either of two main pathways leading to a crossover (CO) or a non-crossover (NCO) recombinant (see Genetic recombination. The pathway leading to a CO involves a double Holliday junction (DHJ) intermediate. Holliday junctions need to be resolved for CO recombination to be completed.
In the budding yeast Saccharomyces cerevisiae , as in the mouse, MLH3 forms a heterodimer with MLH1. Meiotic CO requires resolution of Holliday junctions through actions of the MLH1-MLH3 heterodimer. The MLH1-MLH3 heterodimer is an endonuclease that makes single-strand breaks in supercoiled double-stranded DNA. [9] [10] MLH1-MLH3 binds specifically to Holliday junctions and may act as part of a larger complex to process Holliday junctions during meiosis. [9] MLH1-MLH3 heterodimer (MutL gamma) together with Exo1 and Sgs1 (ortholog of Bloom syndrome helicase) define a joint molecule resolution pathway that produces the majority of crossovers in budding yeast and, by inference, in mammals. [11]

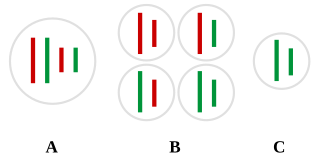
Chromosomal crossover, or crossing over, is the exchange of genetic material during sexual reproduction between two homologous chromosomes' non-sister chromatids that results in recombinant chromosomes. It is one of the final phases of genetic recombination, which occurs in the pachytene stage of prophase I of meiosis during a process called synapsis. Synapsis begins before the synaptonemal complex develops and is not completed until near the end of prophase I. Crossover usually occurs when matching regions on matching chromosomes break and then reconnect to the other chromosome.

Genetic recombination is the exchange of genetic material between different organisms which leads to production of offspring with combinations of traits that differ from those found in either parent. In eukaryotes, genetic recombination during meiosis can lead to a novel set of genetic information that can be further passed on from parents to offspring. Most recombination occurs naturally and can be classified into two types: (1) interchromosomal recombination, occurring through independent assortment of alleles whose loci are on different but homologous chromosomes ; & (2) intrachromosomal recombination, occurring through crossing over.
Gene conversion is the process by which one DNA sequence replaces a homologous sequence such that the sequences become identical after the conversion event. Gene conversion can be either allelic, meaning that one allele of the same gene replaces another allele, or ectopic, meaning that one paralogous DNA sequence converts another.
DNA mismatch repair (MMR) is a system for recognizing and repairing erroneous insertion, deletion, and mis-incorporation of bases that can arise during DNA replication and recombination, as well as repairing some forms of DNA damage.

A Holliday junction is a branched nucleic acid structure that contains four double-stranded arms joined. These arms may adopt one of several conformations depending on buffer salt concentrations and the sequence of nucleobases closest to the junction. The structure is named after Robin Holliday, the molecular biologist who proposed its existence in 1964.

DNA mismatch repair protein Msh2 also known as MutS homolog 2 or MSH2 is a protein that in humans is encoded by the MSH2 gene, which is located on chromosome 2. MSH2 is a tumor suppressor gene and more specifically a caretaker gene that codes for a DNA mismatch repair (MMR) protein, MSH2, which forms a heterodimer with MSH6 to make the human MutSα mismatch repair complex. It also dimerizes with MSH3 to form the MutSβ DNA repair complex. MSH2 is involved in many different forms of DNA repair, including transcription-coupled repair, homologous recombination, and base excision repair.

DNA mismatch repair protein Mlh1 or MutL protein homolog 1 is a protein that in humans is encoded by the MLH1 gene located on chromosome 3. It is a gene commonly associated with hereditary nonpolyposis colorectal cancer. Orthologs of human MLH1 have also been studied in other organisms including mouse and the budding yeast Saccharomyces cerevisiae.

MSH6 or mutS homolog 6 is a gene that codes for DNA mismatch repair protein Msh6 in the budding yeast Saccharomyces cerevisiae. It is the homologue of the human "G/T binding protein," (GTBP) also called p160 or hMSH6. The MSH6 protein is a member of the Mutator S (MutS) family of proteins that are involved in DNA damage repair.

Spo11 is a protein that in humans is encoded by the SPO11 gene. Spo11, in a complex with mTopVIB, creates double strand breaks to initiate meiotic recombination. Its active site contains a tyrosine which ligates and dissociates with DNA to promote break formation. One Spo11 protein is involved per strand of DNA, thus two Spo11 proteins are involved in each double stranded break event.
Chromosome segregation is the process in eukaryotes by which two sister chromatids formed as a consequence of DNA replication, or paired homologous chromosomes, separate from each other and migrate to opposite poles of the nucleus. This segregation process occurs during both mitosis and meiosis. Chromosome segregation also occurs in prokaryotes. However, in contrast to eukaryotic chromosome segregation, replication and segregation are not temporally separated. Instead segregation occurs progressively following replication.

Mismatch repair endonuclease PMS2 is an enzyme that in humans is encoded by the PMS2 gene.

Bloom syndrome protein is a protein that in humans is encoded by the BLM gene and is not expressed in Bloom syndrome.

Exonuclease 1 is an enzyme that in humans is encoded by the EXO1 gene.

PMS1 protein homolog 1 is a protein that in humans is encoded by the PMS1 gene.

MutS protein homolog 5 is a protein that in humans is encoded by the MSH5 gene.

MutS protein homolog 4 is a protein that in humans is encoded by the MSH4 gene.
Crossover junction endonuclease MUS81 is an enzyme that in humans is encoded by the MUS81 gene.

Fanconi anemia, complementation group M, also known as FANCM is a human gene. It is an emerging target in cancer therapy, in particular cancers with specific genetic deficiencies.
Sgs1, also known as slow growth suppressor 1, is a DNA helicase protein found in Saccharomyces cerevisiae. It is a homolog of the bacterial RecQ helicase. Like the other members of the RecQ helicase family, Sgs1 is important for DNA repair. In particular, Sgs1 collaborates with other proteins to repair double-strand breaks during homologous recombination in eukaryotes.

Synthesis-dependent strand annealing (SDSA) is a major mechanism of homology-directed repair of DNA double-strand breaks (DSBs). Although many of the features of SDSA were first suggested in 1976, the double-Holliday junction model proposed in 1983 was favored by many researchers. In 1994, studies of double-strand gap repair in Drosophila were found to be incompatible with the double-Holliday junction model, leading researchers to propose a model they called synthesis-dependent strand annealing. Subsequent studies of meiotic recombination in S. cerevisiae found that non-crossover products appear earlier than double-Holliday junctions or crossover products, challenging the previous notion that both crossover and non-crossover products are produced by double-Holliday junctions and leading the authors to propose that non-crossover products are generated through SDSA.