
A bacteriophage, also known informally as a phage, is a virus that infects and replicates within bacteria and archaea. The term was derived from "bacteria" and the Greek φαγεῖν, meaning "to devour". Bacteriophages are composed of proteins that encapsulate a DNA or RNA genome, and may have structures that are either simple or elaborate. Their genomes may encode as few as four genes and as many as hundreds of genes. Phages replicate within the bacterium following the injection of their genome into its cytoplasm.

Enterobacteria phage λ is a bacterial virus, or bacteriophage, that infects the bacterial species Escherichia coli. It was discovered by Esther Lederberg in 1950. The wild type of this virus has a temperate life cycle that allows it to either reside within the genome of its host through lysogeny or enter into a lytic phase, during which it kills and lyses the cell to produce offspring. Lambda strains, mutated at specific sites, are unable to lysogenize cells; instead, they grow and enter the lytic cycle after superinfecting an already lysogenized cell.
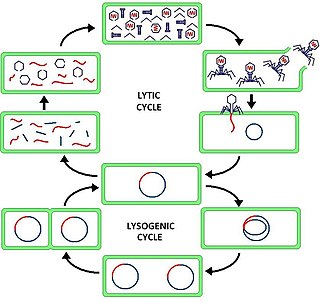
The lytic cycle is one of the two cycles of viral reproduction, the other being the lysogenic cycle. The lytic cycle results in the destruction of the infected cell and its membrane. Bacteriophages that can only go through the lytic cycle are called virulent phages.

Escherichia virus T4 is a species of bacteriophages that infect Escherichia coli bacteria. It is a double-stranded DNA virus in the subfamily Tevenvirinae of the family Straboviridae. T4 is capable of undergoing only a lytic life cycle and not the lysogenic life cycle. The species was formerly named T-even bacteriophage, a name which also encompasses, among other strains, Enterobacteria phage T2, Enterobacteria phage T4 and Enterobacteria phage T6.

Podoviridae was a family of bacteriophage in the order Caudovirales often associated with T-7 like phages. The family and order Caudoviraleshave now been abolished, with the term podovirus now used to refer to the morphology of viruses in this former family. There were 130 species in this family, assigned to 3 subfamilies and 52 genera. This family was characterized by having very short, noncontractile tails. Many former phages in the former family Podoviriade are now classified in the Autographiviridae
Microviridae is a family of bacteriophages with a single-stranded DNA genome. The name of this family is derived from the ancient Greek word μικρός (mikrós), meaning "small". This refers to the size of their genomes, which are among the smallest of the DNA viruses. Enterobacteria, intracellular parasitic bacteria, and spiroplasma serve as natural hosts. There are 22 species in this family, divided among seven genera and two subfamilies.

Viral replication is the formation of biological viruses during the infection process in the target host cells. Viruses must first get into the cell before viral replication can occur. Through the generation of abundant copies of its genome and packaging these copies, the virus continues infecting new hosts. Replication between viruses is greatly varied and depends on the type of genes involved in them. Most DNA viruses assemble in the nucleus while most RNA viruses develop solely in cytoplasm.
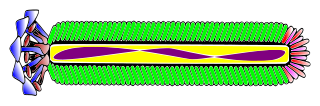
M13 is one of the Ff phages, a member of the family filamentous bacteriophage (inovirus). Ff phages are composed of circular single-stranded DNA (ssDNA), which in the case of the m13 phage is 6407 nucleotides long and is encapsulated in approximately 2700 copies of the major coat protein p8, and capped with about 5 copies each of four different minor coat proteins. The minor coat protein p3 attaches to the receptor at the tip of the F pilus of the host Escherichia coli. The life cycle is relatively short, with the early phage progeny exiting the cell ten minutes after infection. Ff phages are chronic phage, releasing their progeny without killing the host cells. The infection causes turbid plaques in E. coli lawns, of intermediate opacity in comparison to regular lysis plaques. However, a decrease in the rate of cell growth is seen in the infected cells. The replicative form of M13 is circular double-stranded DNA similar to plasmids that are used for many recombinant DNA processes, and the virus has also been used for phage display, directed evolution, nanostructures and nanotechnology applications.

The phi X 174 bacteriophage is a single-stranded DNA (ssDNA) virus that infects Escherichia coli. This virus was isolated in 1935 by Nicolas Bulgakov in Félix d'Hérelle's laboratory at the Pasteur Institute, from samples collected in Paris sewers. Its characterization and the study of its replication mechanism were carried out from the 1950s onwards. It was the first DNA-based genome to be sequenced. This work was completed by Fred Sanger and his team in 1977. In 1962, Walter Fiers and Robert Sinsheimer had already demonstrated the physical, covalently closed circularity of ΦX174 DNA. Nobel prize winner Arthur Kornberg used ΦX174 as a model to first prove that DNA synthesized in a test tube by purified enzymes could produce all the features of a natural virus, ushering in the age of synthetic biology. In 1972–1974, Jerard Hurwitz, Sue Wickner, and Reed Wickner with collaborators identified the genes required to produce the enzymes to catalyze conversion of the single stranded form of the virus to the double stranded replicative form. In 2003, it was reported by Craig Venter's group that the genome of ΦX174 was the first to be completely assembled in vitro from synthesized oligonucleotides. The ΦX174 virus particle has also been successfully assembled in vitro. In 2012, it was shown how its highly overlapping genome can be fully decompressed and still remain functional.
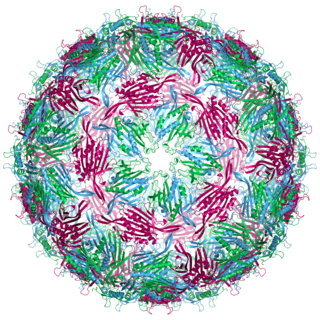
Bacteriophage MS2, commonly called MS2, is an icosahedral, positive-sense single-stranded RNA virus that infects the bacterium Escherichia coli and other members of the Enterobacteriaceae. MS2 is a member of a family of closely related bacterial viruses that includes bacteriophage f2, bacteriophage Qβ, R17, and GA.
P1 is a temperate bacteriophage that infects Escherichia coli and some other bacteria. When undergoing a lysogenic cycle the phage genome exists as a plasmid in the bacterium unlike other phages that integrate into the host DNA. P1 has an icosahedral head containing the DNA attached to a contractile tail with six tail fibers. The P1 phage has gained research interest because it can be used to transfer DNA from one bacterial cell to another in a process known as transduction. As it replicates during its lytic cycle it captures fragments of the host chromosome. If the resulting viral particles are used to infect a different host the captured DNA fragments can be integrated into the new host's genome. This method of in vivo genetic engineering was widely used for many years and is still used today, though to a lesser extent. P1 can also be used to create the P1-derived artificial chromosome cloning vector which can carry relatively large fragments of DNA. P1 encodes a site-specific recombinase, Cre, that is widely used to carry out cell-specific or time-specific DNA recombination by flanking the target DNA with loxP sites.

T7 DNA polymerase is an enzyme used during the DNA replication of the T7 bacteriophage. During this process, the DNA polymerase “reads” existing DNA strands and creates two new strands that match the existing ones. The T7 DNA polymerase requires a host factor, E. coli thioredoxin, in order to carry out its function. This helps stabilize the binding of the necessary protein to the primer-template to improve processivity by more than 100-fold, which is a feature unique to this enzyme. It is a member of the Family A DNA polymerases, which include E. coli DNA polymerase I and Taq DNA polymerase.

Bacteriophage Qbeta, commonly referred to as Qbeta or Qβ, is a species consisting of several strains of positive-strand RNA virus which infects bacteria that have F-pili, most commonly Escherichia coli. Its linear genome is packaged into an icosahedral capsid with a diameter of 28 nm. Bacteriophage Qβ enters its host cell after binding to the side of the F-pilus.
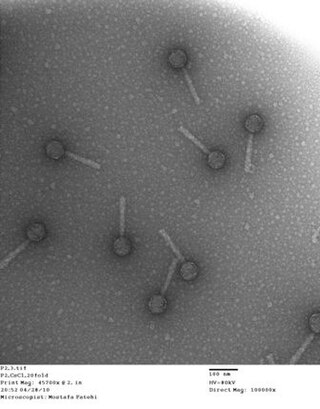
Bacteriophage P2, scientific name Peduovirus P2, is a temperate phage that infects E. coli. It is a tailed virus with a contractile sheath and is thus classified in the genus Peduovirus, family Peduoviridae within class Caudoviricetes. This genus of viruses includes many P2-like phages as well as the satellite phage P4.
Phikmvvirus is a genus of viruses that infect bacteria. There are currently 16 species in this genus including the type species Pseudomonas virus phiKMV. Bacteriophage phiKMV and its relatives are known to be highly virulent phages, producing large clear plaques on a susceptible host. The only reported exception is phage LKA1, which yields small plaques surrounded by a halo. While all other P. aeruginosa-specific phikmvviruses use the Type IV pili as primary receptor, LKA1 particles attach to the bacterial lipopolysaccharide layer.
Enquatrovirus is a genus of bacteriophages in the order Caudovirales, in the family Podoviridae. Bacteria serve as natural hosts. There is currently only one species in this genus: the type species Escherichia virus N4.
Escherichia virus CC31, formerly known as Enterobacter virus CC31, is a dsDNA bacteriophage of the subfamily Tevenvirinae responsible for infecting the bacteria family of Enterobacteriaceae. It is one of two discovered viruses of the genus Karamvirus, diverging away from the previously discovered T4virus, as a clonal complex (CC). CC31 was first isolated from Escherichia coli B strain S/6/4 and is primarily associated with Escherichia, even though is named after Enterobacter.

Duplodnaviria is a realm of viruses that includes all double-stranded DNA viruses that encode the HK97 fold major capsid protein. The HK97 fold major capsid protein is the primary component of the viral capsid, which stores the viral deoxyribonucleic acid (DNA). Viruses in the realm also share a number of other characteristics, such as an icosahedral capsid, an opening in the viral capsid called a portal, a protease enzyme that empties the inside of the capsid prior to DNA packaging, and a terminase enzyme that packages viral DNA into the capsid.
Bacteriophage AP205 is a plaque-forming bacteriophage that infects Acinetobacter bacteria. Bacteriophage AP205 is a protein-coated virus with a positive single-stranded RNA genome. It is a member of the family Fiersviridae, consisting of particles that infect Gram-negative bacteria such as E. coli.
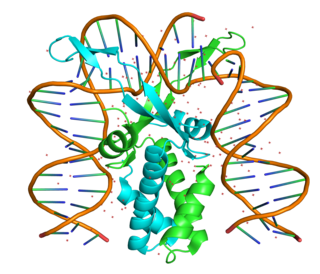
The integration host factor (IHF) is a bacterial DNA binding protein complex that facilitates genetic recombination, replication, and transcription by binding to specific DNA sequences and bending the DNA. It also facilitates the integration of foreign DNA into the host genome. It is a heterodimeric complex composed of two homologous subunits IHFalpha and IHFbeta.






















