
The structural formula of a chemical compound is a graphic representation of the molecular structure, showing how the atoms are possibly arranged in the real three-dimensional space. The chemical bonding within the molecule is also shown, either explicitly or implicitly. Unlike other chemical formula types, which have a limited number of symbols and are capable of only limited descriptive power, structural formulas provide a more complete geometric representation of the molecular structure. For example, many chemical compounds exist in different isomeric forms, which have different enantiomeric structures but the same molecular formula. There are multiple types of ways to draw these structural formulas such as: Lewis structures, condensed formulas, skeletal formulas, Newman projections, Cyclohexane conformations, Haworth projections, and Fischer projections.

Molecular dynamics (MD) is a computer simulation method for analyzing the physical movements of atoms and molecules. The atoms and molecules are allowed to interact for a fixed period of time, giving a view of the dynamic "evolution" of the system. In the most common version, the trajectories of atoms and molecules are determined by numerically solving Newton's equations of motion for a system of interacting particles, where forces between the particles and their potential energies are often calculated using interatomic potentials or molecular mechanical force fields. The method is applied mostly in chemical physics, materials science, and biophysics.
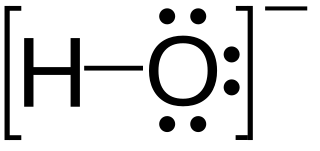
In chemistry, a lone pair refers to a pair of valence electrons that are not shared with another atom in a covalent bond and is sometimes called an unshared pair or non-bonding pair. Lone pairs are found in the outermost electron shell of atoms. They can be identified by using a Lewis structure. Electron pairs are therefore considered lone pairs if two electrons are paired but are not used in chemical bonding. Thus, the number of electrons in lone pairs plus the number of electrons in bonds equals the number of valence electrons around an atom.
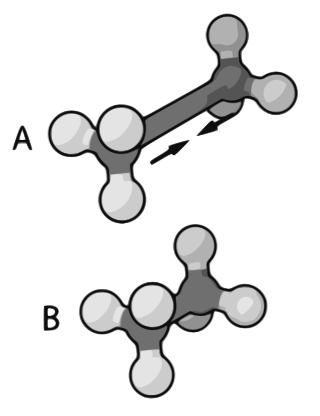
Molecular mechanics uses classical mechanics to model molecular systems. The Born–Oppenheimer approximation is assumed valid and the potential energy of all systems is calculated as a function of the nuclear coordinates using force fields. Molecular mechanics can be used to study molecule systems ranging in size and complexity from small to large biological systems or material assemblies with many thousands to millions of atoms.

The skeletal formula, line-angle formula, bond-line formula or shorthand formula of an organic compound is a type of molecular structural formula that serves as a shorthand representation of a molecule's bonding and some details of its molecular geometry. A skeletal formula shows the skeletal structure or skeleton of a molecule, which is composed of the skeletal atoms that make up the molecule. It is represented in two dimensions, as on a piece of paper. It employs certain conventions to represent carbon and hydrogen atoms, which are the most common in organic chemistry.

Molecular modelling encompasses all methods, theoretical and computational, used to model or mimic the behaviour of molecules. The methods are used in the fields of computational chemistry, drug design, computational biology and materials science to study molecular systems ranging from small chemical systems to large biological molecules and material assemblies. The simplest calculations can be performed by hand, but inevitably computers are required to perform molecular modelling of any reasonably sized system. The common feature of molecular modelling methods is the atomistic level description of the molecular systems. This may include treating atoms as the smallest individual unit, or explicitly modelling protons and neutrons with its quarks, anti-quarks and gluons and electrons with its photons.
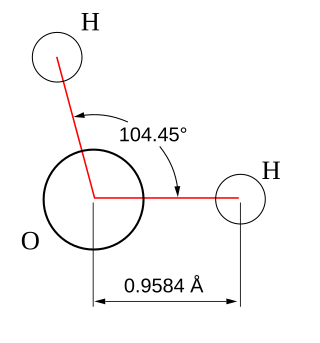
Molecular geometry is the three-dimensional arrangement of the atoms that constitute a molecule. It includes the general shape of the molecule as well as bond lengths, bond angles, torsional angles and any other geometrical parameters that determine the position of each atom.
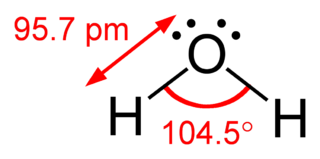
Valence shell electron pair repulsion (VSEPR) theory is a model used in chemistry to predict the geometry of individual molecules from the number of electron pairs surrounding their central atoms. It is also named the Gillespie-Nyholm theory after its two main developers, Ronald Gillespie and Ronald Nyholm.
In chemistry, orbital hybridisation is the concept of mixing atomic orbitals to form new hybrid orbitals suitable for the pairing of electrons to form chemical bonds in valence bond theory. For example, in a carbon atom which forms four single bonds, the valence-shell s orbital combines with three valence-shell p orbitals to form four equivalent sp3 mixtures in a tetrahedral arrangement around the carbon to bond to four different atoms. Hybrid orbitals are useful in the explanation of molecular geometry and atomic bonding properties and are symmetrically disposed in space. Usually hybrid orbitals are formed by mixing atomic orbitals of comparable energies.
A tetrahedral intermediate is a reaction intermediate in which the bond arrangement around an initially double-bonded carbon atom has been transformed from trigonal to tetrahedral. Tetrahedral intermediates result from nucleophilic addition to a carbonyl group. The stability of tetrahedral intermediate depends on the ability of the groups attached to the new tetrahedral carbon atom to leave with the negative charge. Tetrahedral intermediates are very significant in organic syntheses and biological systems as a key intermediate in esterification, transesterification, ester hydrolysis, formation and hydrolysis of amides and peptides, hydride reductions, and other chemical reactions.
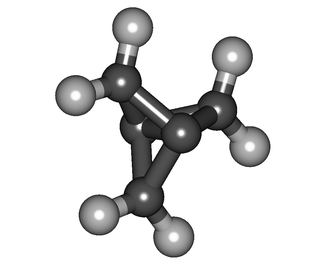
In organic chemistry, ring strain is a type of instability that exists when bonds in a molecule form angles that are abnormal. Strain is most commonly discussed for small rings such as cyclopropanes and cyclobutanes, whose internal angles are substantially smaller than the idealized value of approximately 109°. Because of their high strain, the heat of combustion for these small rings is elevated.
In quantum chemistry, the quantum theory of atoms in molecules (QTAIM), sometimes referred to as atoms in molecules (AIM), is a model of molecular and condensed matter electronic systems in which the principal objects of molecular structure - atoms and bonds - are natural expressions of a system's observable electron density distribution function. An electron density distribution of a molecule is a probability distribution that describes the average manner in which the electronic charge is distributed throughout real space in the attractive field exerted by the nuclei. According to QTAIM, molecular structure is revealed by the stationary points of the electron density together with the gradient paths of the electron density that originate and terminate at these points.

In the context of chemistry, molecular physics, physical chemistry, and molecular modelling, a force field is a computational model that is used to describe the forces between atoms within molecules or between molecules as well as in crystals. Force fields are a variety of interatomic potentials. More precisely, the force field refers to the functional form and parameter sets used to calculate the potential energy of a system on the atomistic level. Force fields are usually used in molecular dynamics or Monte Carlo simulations. The parameters for a chosen energy function may be derived from classical laboratory experiment data, calculations in quantum mechanics, or both. Force fields utilize the same concept as force fields in classical physics, with the main difference being that the force field parameters in chemistry describe the energy landscape on the atomistic level. From a force field, the acting forces on every particle are derived as a gradient of the potential energy with respect to the particle coordinates.
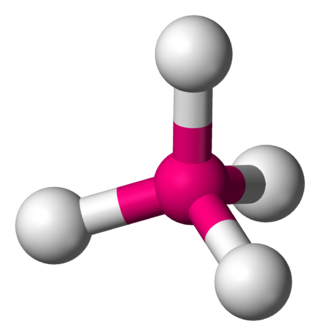
In a tetrahedral molecular geometry, a central atom is located at the center with four substituents that are located at the corners of a tetrahedron. The bond angles are cos−1(−1⁄3) = 109.4712206...° ≈ 109.5° when all four substituents are the same, as in methane as well as its heavier analogues. Methane and other perfectly symmetrical tetrahedral molecules belong to point group Td, but most tetrahedral molecules have lower symmetry. Tetrahedral molecules can be chiral.
Molecular graphics is the discipline and philosophy of studying molecules and their properties through graphical representation. IUPAC limits the definition to representations on a "graphical display device". Ever since Dalton's atoms and Kekulé's benzene, there has been a rich history of hand-drawn atoms and molecules, and these representations have had an important influence on modern molecular graphics.
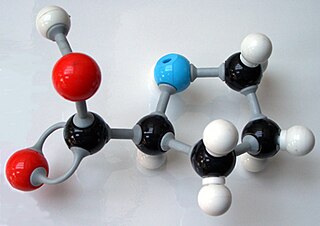
In chemistry, the ball-and-stick model is a molecular model of a chemical substance which displays both the three-dimensional position of the atoms and the bonds between them. The atoms are typically represented by spheres, connected by rods which represent the bonds. Double and triple bonds are usually represented by two or three curved rods, respectively, or alternately by correctly positioned sticks for the sigma and pi bonds. In a good model, the angles between the rods should be the same as the angles between the bonds, and the distances between the centers of the spheres should be proportional to the distances between the corresponding atomic nuclei. The chemical element of each atom is often indicated by the sphere's color.
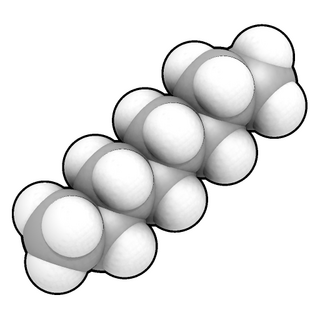
In chemistry, a space-filling model, also known as a calotte model, is a type of three-dimensional (3D) molecular model where the atoms are represented by spheres whose radii are proportional to the radii of the atoms and whose center-to-center distances are proportional to the distances between the atomic nuclei, all in the same scale. Atoms of different chemical elements are usually represented by spheres of different colors.
In chemistry, the Z-matrix is a way to represent a system built of atoms. A Z-matrix is also known as an internal coordinate representation. It provides a description of each atom in a molecule in terms of its atomic number, bond length, bond angle, and dihedral angle, the so-called internal coordinates, although it is not always the case that a Z-matrix will give information regarding bonding since the matrix itself is based on a series of vectors describing atomic orientations in space. However, it is convenient to write a Z-matrix in terms of bond lengths, angles, and dihedrals since this will preserve the actual bonding characteristics. The name arises because the Z-matrix assigns the second atom along the Z axis from the first atom, which is at the origin.
The van der Waals surface of a molecule is an abstract representation or model of that molecule, illustrating where, in very rough terms, a surface might reside for the molecule based on the hard cutoffs of van der Waals radii for individual atoms, and it represents a surface through which the molecule might be conceived as interacting with other molecules. Also referred to as a van der Waals envelope, the van der Waals surface is named for Johannes Diderik van der Waals, a Dutch theoretical physicist and thermodynamicist who developed theory to provide a liquid-gas equation of state that accounted for the non-zero volume of atoms and molecules, and on their exhibiting an attractive force when they interacted. van der Waals surfaces are therefore a tool used in the abstract representations of molecules, whether accessed, as they were originally, via hand calculation, or via physical wood/plastic models, or now digitally, via computational chemistry software. Practically speaking, CPK models, developed by and named for Robert Corey, Linus Pauling, and Walter Koltun, were the first widely used physical molecular models based on van der Waals radii, and allowed broad pedagogical and research use of a model showing the van der Waals surfaces of molecules.
In chemistry, a halogen bond occurs when there is evidence of a net attractive interaction between an electrophilic region associated with a halogen atom in a molecular entity and a nucleophilic region in another, or the same, molecular entity. Like a hydrogen bond, the result is not a formal chemical bond, but rather a strong electrostatic attraction. Mathematically, the interaction can be decomposed in two terms: one describing an electrostatic, orbital-mixing charge-transfer and another describing electron-cloud dispersion. Halogen bonds find application in supramolecular chemistry; drug design and biochemistry; crystal engineering and liquid crystals; and organic catalysis.


















