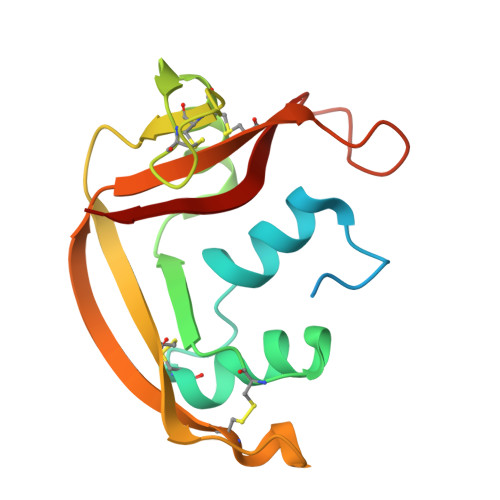
Triazole double-headed ribonucleosides as inhibitors of eosinophil derived neurotoxin.
Chatzileontiadou, D.S., Parmenopoulou, V., Manta, S., Kantsadi, A.L., Kylindri, P., Griniezaki, M., Kontopoulou, F., Telopoulou, A., Prokova, H., Panagopoulos, D., Boix, E., Balatsos, N.A., Komiotis, D., Leonidas, D.D.(2015) Bioorg Chem 63: 152-165
- PubMed: 26551065
- DOI: https://doi.org/10.1016/j.bioorg.2015.10.007
- Primary Citation of Related Structures:
5E13 - PubMed Abstract:
Eosinophil derived neurotoxin (EDN) is an eosinophil secretion protein and a member of the Ribonuclease A (RNase A) superfamily involved in the immune response system and inflammatory disorders. The pathological actions of EDN are strongly dependent on the enzymatic activity and therefore, it is of significant interest to discover potent and specific inhibitors of EDN. In this framework we have assessed the inhibitory potency of triazole double-headed ribonucleosides. We present here an efficient method for the heterologous production and purification of EDN together with the synthesis of nucleosides and their biochemical evaluation in RNase A and EDN. Two groups of double-headed nucleosides were synthesized by the attachment of a purine or a pyrimidine base, through a triazole group at the 3'-C position of a pyrimidine or a purine ribonucleoside, respectively. Based on previous data with mononucleosides these compounds were expected to improve the inhibitory potency for RNase A and specificity for EDN. Kinetics data revealed that despite the rational, all but one, double-headed ribonucleosides were less potent than the respective mononucleosides while they were also more specific for ribonuclease A than for EDN. Compound 11c (9-[3'-[4-[(cytosine-1-yl)methyl]-1,2,3-triazol-1-yl]-β-d-ribofuranosyl]adenine) displayed a stronger preference for EDN than for ribonuclease A and a Ki value of 58μM. This is the first time that an inhibitor is reported to have a better potency for EDN than for RNase A. The crystal structure of EDN-11c complex reveals the structural basis of its potency and selectivity providing important guidelines for future structure-based inhibitor design efforts.
Organizational Affiliation:
Department of Biochemistry and Biotechnology, University of Thessaly, 26 Ploutonos Str., 41221 Larissa, Greece.